SAM-TB
Home
My data
My analysis
My samples
Help
FAQ
中文
Sign in
##Single sample variants analysis ###Operating instructions (1) Create analysis: for samples whose sequencing data have been completely uploaded, the “Single sample variants analysis” can be started in the following two ways: ① Create analysis for a single sample: click “+ New analysis” and pull-down menu select “Single sample variants analysis”. Click “Select sample” to select a sample, or click “New sample” to create a sample. Then click “Modifiable Parameters” to modify the analysis parameters and then click “Start” to begin the analysis. If the parameters are not modified, the default parameters will be used for analysis. 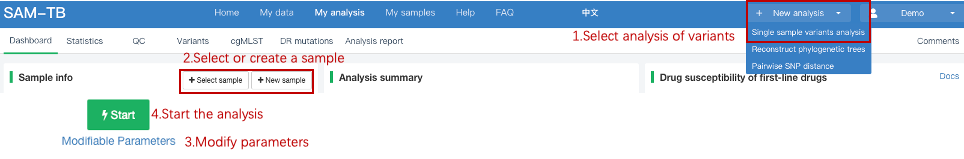 Note: to create an analysis for a new sample, the sequencing data corresponding to the sample should be added or uploaded (if not previously uploaded). 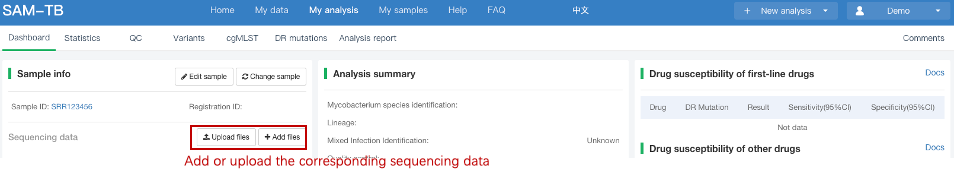 ② Create analyses for multiple samples: click the person avatar in the upper right corner of the platform and select “Analysis setting”. Modify the analysis and quality control parameters on the webpage that appears. Click “My data”, then click “Create batch analysis” in the page. In the floating window, click "Download sample import template" to download the template file. Fill in sample information according to the template and upload the edited file to the website. If the upload is successful, the website will automatically perform "Single sample variants analysis" for all samples listed in the file. If the parameters are not modified, the default parameters will be used for analysis. 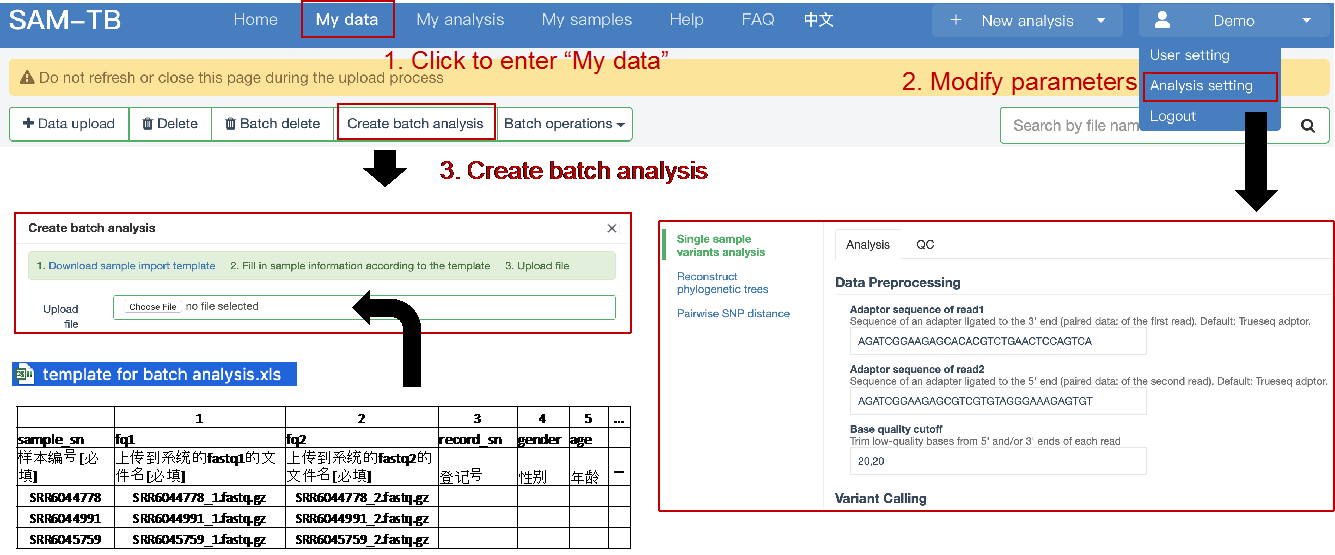 (2) To stop the analysis: the analyses with status shown as “Running” can be stopped in the two ways: ① Tick the box next to the samples, click “Batch operations”, and select “Batch terminate” to stop the selected analyses. ② Click on the ID of the sample to enter the “Dashboard” page, then click “Stop” at the bottom of the page to stop the analysis. 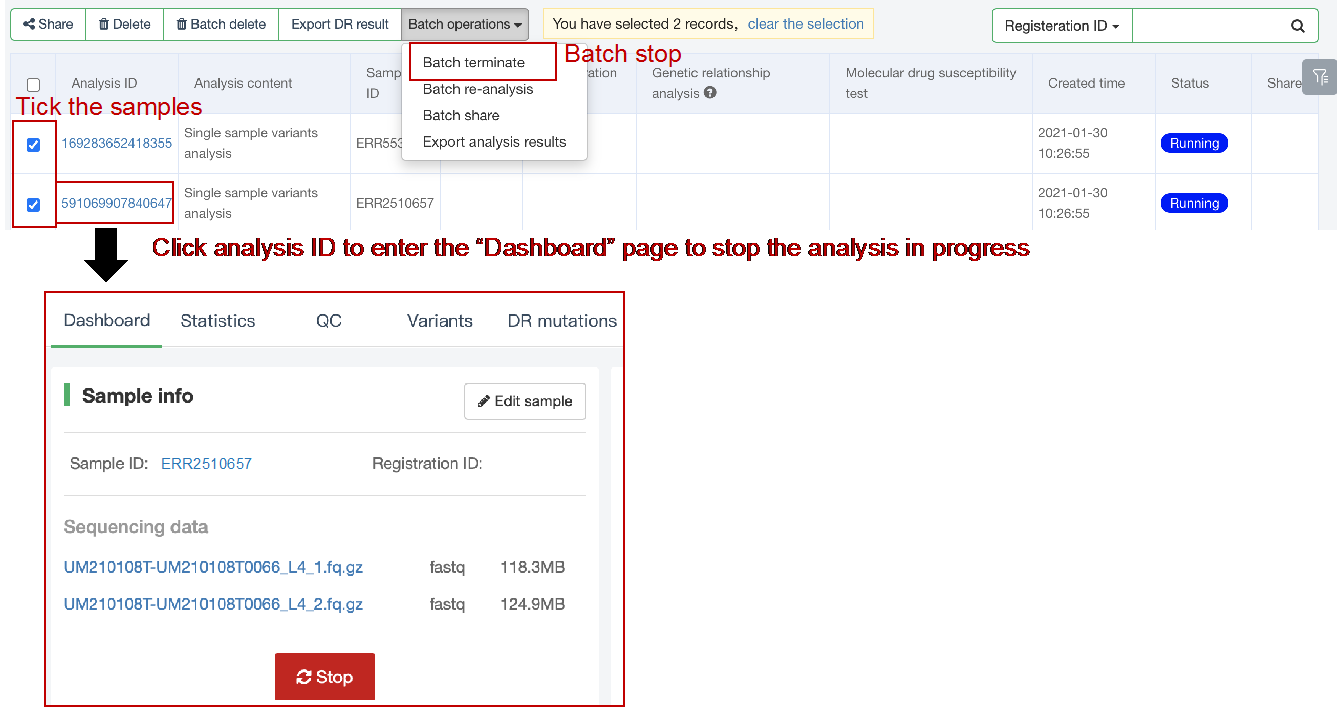 (3) Re-analysis: The analyses with “Error” status can be reanalyzed in two ways: ① Click the person avatar in the upper right corner of the platform and select “Analysis setting”. Modify the analysis and quality control parameters in the page that appears (refers to “Create analyses for multiple samples”). Return to the webpage of “My analysis”. Tick the box to the left of the samples, click “Batch operations”, and select “Batch re-analysis” to reanalyze selected ones. ② Click the ID of a sample that has been analyzed to enter the “Dashboard” page, then click “Modifiable Parameters” at the bottom to modify the parameters as desired, and then click “Restart” to reanalyze. Note: You can also use the default parameters for re-analysis. Any reanalysis will overwrite results of the previous analysis. 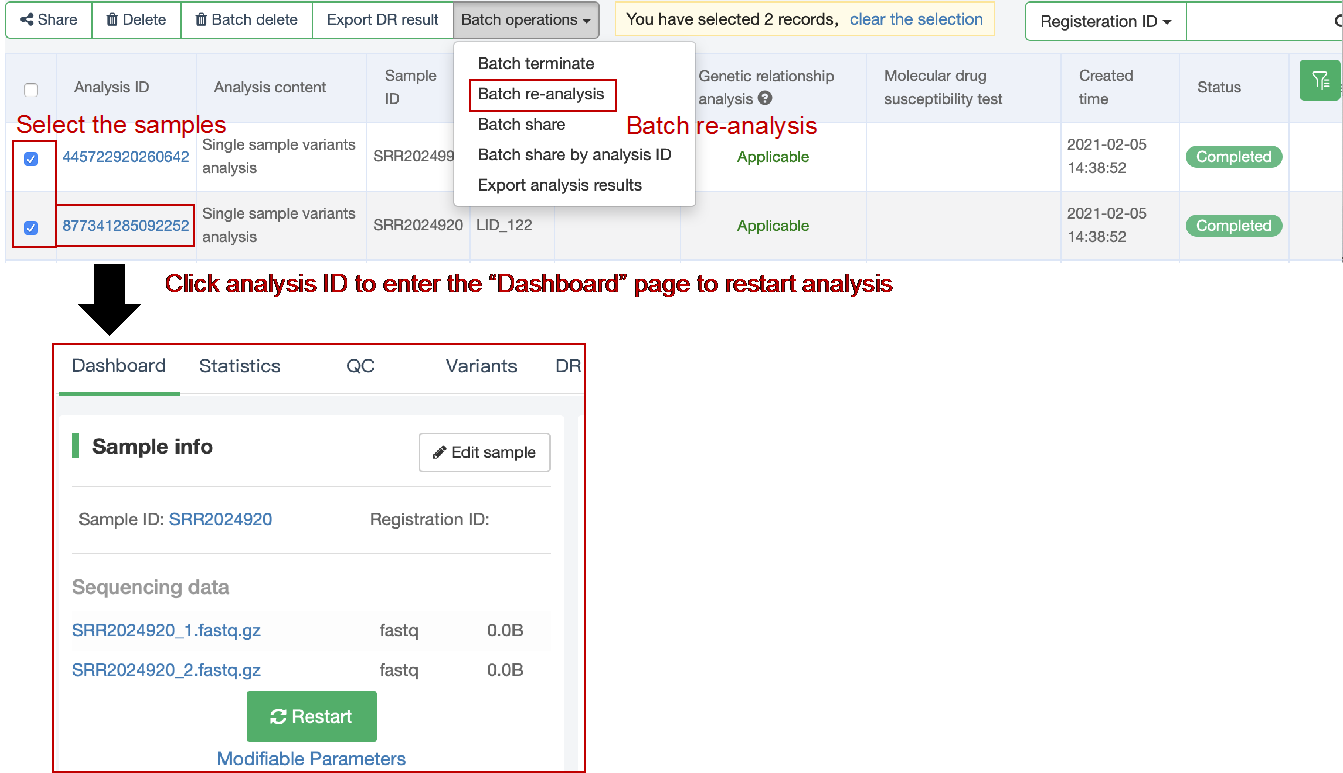 ###Explanation of results After the analysis is completed normally, click the analysis ID to view its analysis results. You can switch to different result pages by clicking on “Dashboard”, “Statistics”, “QC”, “Variants”, “cgMLST”, “DR mutations” and “Analysis report” in the secondary navigation bar.  (1) The “Dashboard” page shows the analysis log and the main reaults. 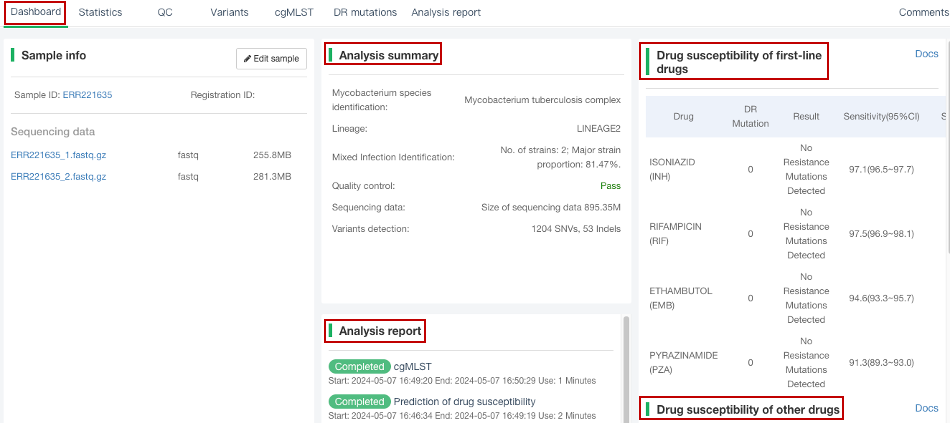 (2) The "Statistics" page details the results of sequencing quality, sequencing depth, the number of detected mutations, etc. 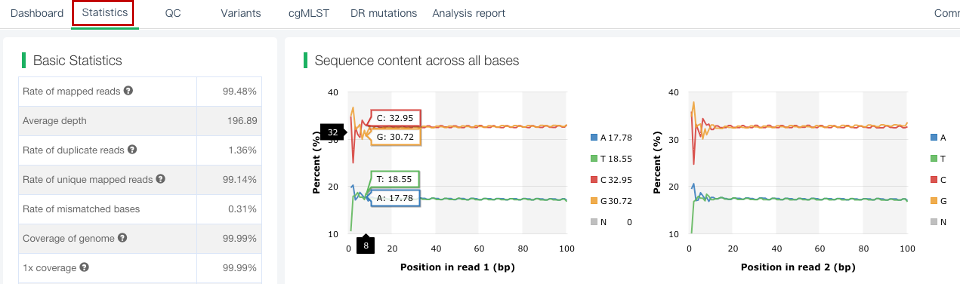 (3) The "QC" page shows the quality control results, including the quality of the sequencing data and the results of genome mapping to the reference sequence (H37Rv, NC_000962.2). 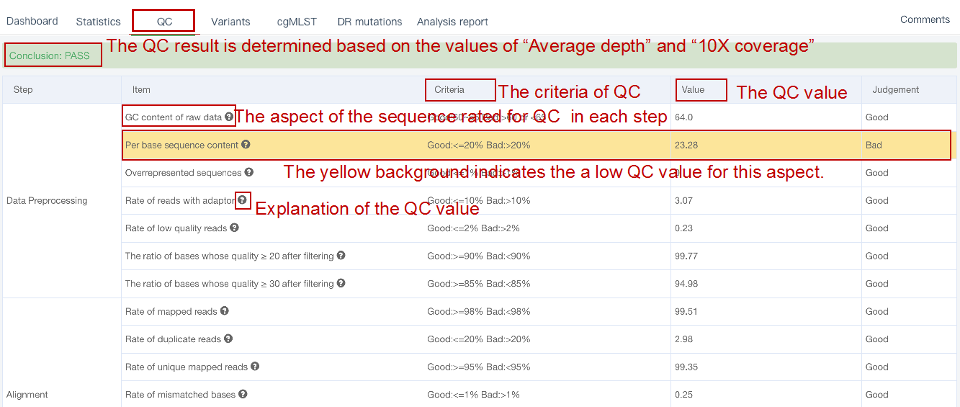 Note: The website uses the values of “Average depth” and “10X coverage” to determine whether the result of quality control is pass or failed. However, if the quality value of multiple items in the "Preprocessing" step are judged “bad”, the sequencing quality of the sample is poor. The poor sequencing quality may affect the prediction of sensitivity to first-line drugs, even though the overall quality control of the sample is passed. Similarly, if multiple values in the “Alignment” steps are bad, the prediction of sensitivity to first-line drugs may also be affected. (4) The "Variants" page shows the results of the genome-wide variants detection. Click “Export” to export all variants and their annotations. Enter a gene name in the search box to search for all variants in a particular gene. Click the screening icon on right side of the page to select or reorder the variants. 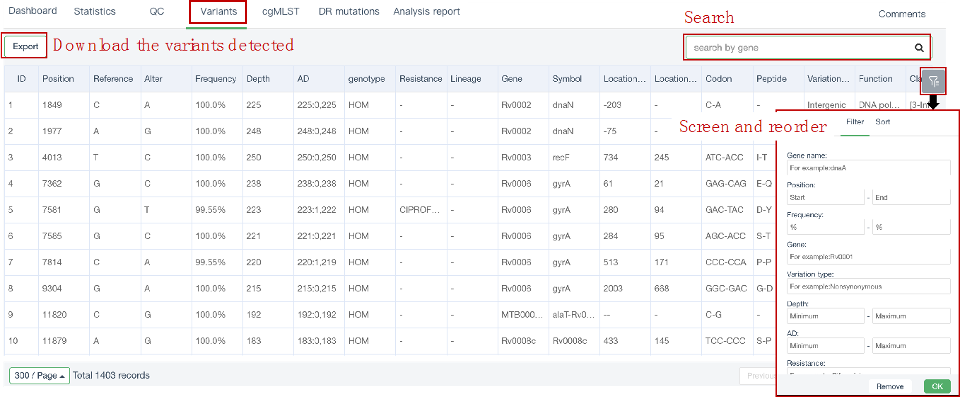 (5) The “cgMLST” page shows the cgMLST allelic profile of the sample. Click “Export” to export the genotypes corresponding to all cgMLST genes. Enter a gene name in the search box to search for genotypes in a particular gene. Click the screening icon on right side of the page to perform selecting or reordering. 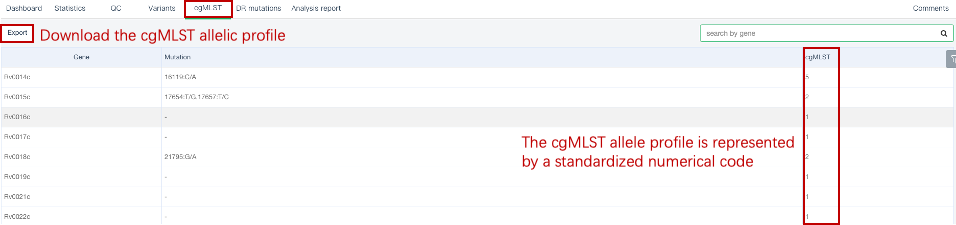 (6) The “DR mutations” page shows drug resistance mutations. Click “Export” to export all drug resistance mutations. Enter a gene name in the search box to search for drug resistance mutations on a particular gene. Click the screening icon on the right side of the page to screen or reorder the drug resistance mutations. (7) The “Analysis report” page shows the results of “single sample variants analysis”, including the results of species/lineage identification, mixed infections identification, cgMLST analysis, drug resistance prediction and drug resistance mutations, etc. Click “Download” icon in the upper right corner to download the word version of the report.
<< Return
Title:
Description:
Thank you for using our service, we will reply you by email as soon as possible.